|
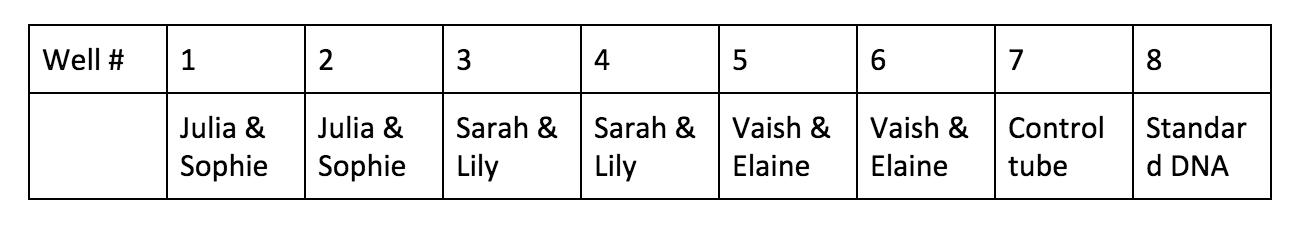
Lab partner: Vaishnavi Kumar Introduction: PCR, short for polymerase chain reaction, is a useful way to help DNA scientists to amplify DNA segments and better understand it. Through PCR, we can make millions of copies of a specific region of DNA. So PCR is basically a special kind of DNA replication, but in a test tube!! Imagine if you have some hair or blood from a crime scene years ago, even the amount is minimal, as long as you can separate out a segment of DNA, then using PCR, you can “amplify” them and compare to match the criminals. In this lab, we will use enzyme Taq polymerase, which is very stable at high temperature. The results of the polymerase chain reaction are visualized using gel electrophoresis, the technique we practiced two weeks ago. Materials and Methods: Material list: 1% buffer TBE Gel tray Gel rig with comb 1 roll electrical tape Gloves Eppendorf tubes Micropipettor Thermocycler Enzyme “bead” Ice tank 10µL λ DNA Centrifuge Procedure Day 1 01.30.18 F Tue. Cast the gel and PCR We use 1% buffer TBE to cast the gel, and then carefully put them in a Ziploc bag and store in the fridge. Be sure to put on gloves for the PCR process, the oil on our hands can cause contamination which can affect the final results. We pair up into 3 groups of 2. For each big group, we have a control tube, which contains all the same contents, but unlike other test tubes, it doesn’t go through the thermocycler. We assign the well # to each pair. Obtain the 2 PCR tubes. We notice that there’s already substance inside the tube, which contains enzyme Taq polymerase, nucleotides and some salts needed for a successful reaction. Label each test tube, then put them into the ice holder.
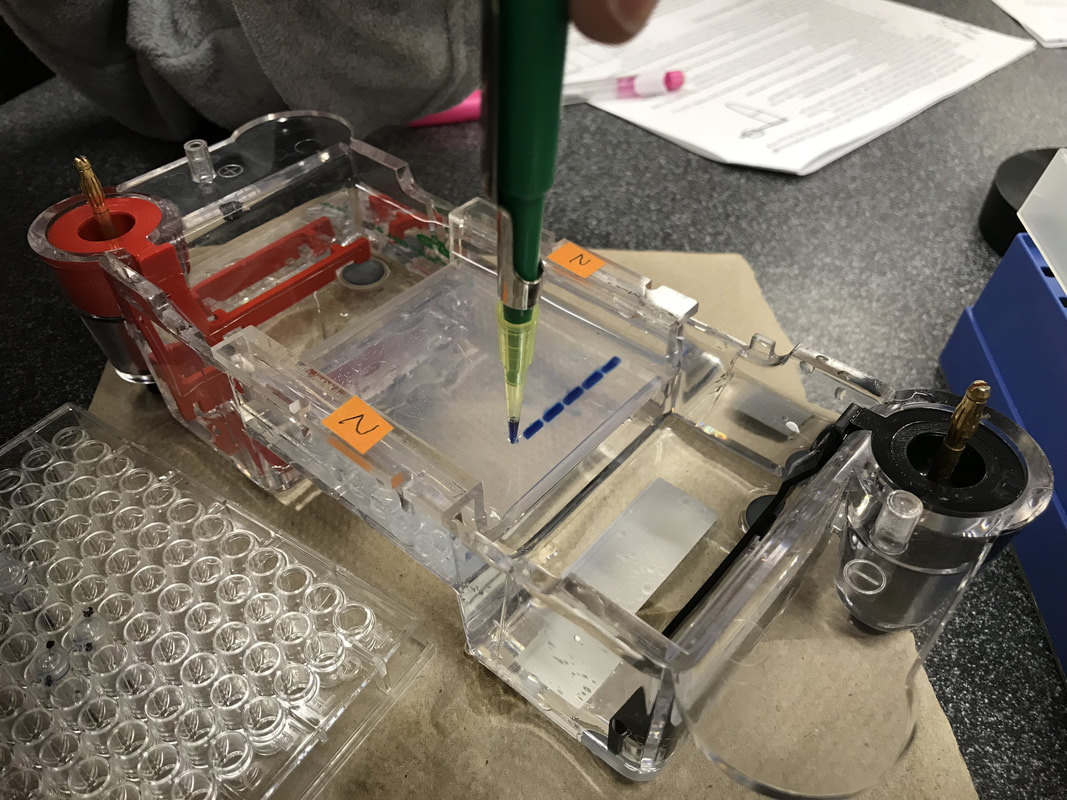
Procedure Day 2 02.02.18 C Fri. Gel electrophoresis, visualize, and analyze the results: Add 4μL of UView loading dye to the PCR test tubes and the control tube. So after we finish the gel electrophoresis, we can be able to observe the DNA bands under the UV light. Analyze the gel over UV light, here’s a photo of our results! Results: The distance traveled by each DNA bands is measured in centimeter. Be careful when measuring the standard DNA, which has has six fragments. * : an error occurred during the experiment. The base pairs # of each of the six bands in the standard DNA is given by the teacher. Plot the point on a logarithmic scale graph. Plot the curve according to the coordinates from known distance migrated and bas pairs # of the standard DNA. The line and dots in red means estimated values. Predict/estimate the number of base pairs(y) using distance traveled(x) by extending the curve. * : an error occurred during the experiment. Conclusion: In this lab, we learned about how to perform PCR(polymerase chain reaction) using λ DNA and what happened in the tiny test tube and the control test tube. After this lab, I realize the whole experiment is mainly divided into two parts. First, the polymerase chain reaction with the help of thermocycler. Second, a visualized version of what the reaction looks like using the gel electrophoresis and UV dye helps to let our eyes better see and analyze them. For the reaction to successfully happen, it’s crucial for the test tubes to go through thermocycler because the DNA had to denature at the right temperature and the enzyme polymerase works the best under certain lower temperature. To prove this, we conduct a control tube(well #7) which doesn’t go through temperature changes in the thermocycler. From the results picture above, it’s clear that in all the successfully reactions(well #3-6), DNA bands traveled around 3.20-3.25cm. However, the mixture in control tube(well #7)doesn’t travel that far although it has all the contents needed for the reaction to happen. The control test tube(well #7) teaches us the importance of going through the right temperatures, and the error occurred with Julia and Sophie’s test tubes(well #1&2) also shows us the importance of the presence of BOTH the anneal primers and the λ DNA. According to Julia and Sophie, they misread the instructions and added only the anneal primers into their first test tube(well #1) and only the λ DNA into their second test tube(well #2). Though both test tubes had the Taq enzymes, nucleotides, and other chemicals required for a successful reaction, and went through the temperature cycles in the thermocycler, the results are not positive. The first test tube with only the anneal primer(well #1) appears on the final gel results but didn’t really move; the second test tube with only the λ DNA didn’t even appear on the final gel picture. To avoid this type of experimental error, be sure to read the instructions diligently and make sure to understand the importance of the presence of all the chemicals. Below are several questions that may help guide me for further investigation: What is crucial about Taq polymerase to the PCR? What does Taq polymerase make out of? Why makes it stable at high temperature? What does each part of “denature - anneal - extend” means? What do the Taq polymerase, the primer, and the λ DNA look like in each of the stages? What does it mean by “anneal”? What substance “extends”? Discussion: PCR, polymerase chain reaction, is the repetition of the cycle “denature - anneal - extend” over and over again in order to make millions of copies of DNA in short time and amplify a certain region of DNA so we can better analyze and understand it. The first step, denature, uses the special feature that DNA has at high temperature: it denatures and becomes single-stranded at around 95℃. Then the second step, anneal, the temperature is readjusted to where the primer will best react with DNA, which is around 55-60℃. Thirdly, extend, the Taq polymerase extends the primer at 5’-3’ direction and duplicates the DNA fragments between primers. The Taq polymerase will “synthesizes -- builds -- two new stranded DNA, using the original strands as templates”(NHGRI). Then each of these new strands can be used to create two new copies, and so on. The cycle of denaturing and synthesizing new DNA is repeated, sometimes “as many as 30 or 40 times, leading to more than one billion exact copies of the original DNA segment”(NHGRI). From the picture of gel, we can tell the significant difference between test tubes that were in the thermocycler(well #3-6 at around 3.20cm, 1540-1570 base pairs) and the control tube that wasn’t(well #7 at around 1.22cm, 23,130 base pairs). Without putting the control test tube into the thermocycler, the mixture in that tube did NOT go through any of the denature - anneal - extend cycle at all. Though inside the control tube has all the contents needed for the reaction, such as Taq polymerase, primers, and λ DNA, the chain reaction can’t be performed without going through the right temperatures for the “denature - anneal - extend” cycle. Polymerase Chaine Reaction: Image credit: https://www.genome.gov/images/content/pcr_factsheet.jpg DNA replication process in the cell and same process in the test tubes in PCR share resemblances, but there are also differences. Firstly, to unzip the double strands into a single strand. In DNA replication in the cell, helicase unzips the double-stranded molecule. However, it’s costly to produce. In PCR, we use another solution to unzip the DNA: heat denature at around 95℃. Secondly, the RNA primer, a signal needed for DNA polymerase to start combining nucleotides onto the single strand inside an organism’s cell. In PCR, thousands of specially designed primers can anneal to the DNA when the temperature is cooling down from previous denaturing step. Thirdly, the polymerase. In the cell replication, DNA polymerase is responsible keep combining nucleotides and extend the growing DNA strand. In PCR, we still use DNA polymerase, but a special one called Taq DNA polymerase. Thermus Aquaticus. Picture credit: https://en.wikipedia.org/wiki/Thermus_aquaticus#/media/File:Thermus_aquaticus.JPG Taq polymerase comes from the Thermus aquaticus, a species of bacteria that live in hot spring and boiling mud. Taq polymerase derived from these bacteria is a milestone discovering. Because like the bacteria in its name, Taq DNA polymerase is very thermostable even at the high temperature when DNA denatures. Other polymerase enzyme derived from some other bacterium would have already denatured during the process of unzipping DNA. It can also be used repeatedly around 25-30 times before it breaks down. Taq polymerase not only stay stable under high temperature, it also makes the PCR much more efficient and cost-effective. DNA is naturally colorless, so it must be stained so that we can better visualize the DNA bands and their locations. The UV dye we used makes the DNA bands glow under the UV light, which is much easier for us to do all the measuring and analysis. Some “Tracking dye which also contains a component(usually glycerol or sucrose) to increase the density of the sample to facilitate the loading”(bioinformatics). The known size of the amplicon is 1,106 base pairs given by our instructor, which is around 400 base pairs fewer than our closest results. There are a few possible inaccuracies which can lead to bigger experimental # of base pairs than the actual value. Since we plot the graph on a logarithmic scale by hands, and we have to estimate the extended graph, it’s very likely that the point deviates the best fit line. Extended BONUS question: Use the following information to calculate how many target DNA molecules were present in the 4.0 nanograms of λ DNA that you added to your PCR reaction:
Works cited and reference: "Polymerase Chain Reaction (PCR) Fact Sheet." National Human Genome Research Institute (NHGRI), 2015, https://www.genome.gov/10000207/polymerase-chain-reaction-pcr-fact-sheet/#al-1.
"PCR." Sci.Sdsu.Edu, 2002, http://www.sci.sdsu.edu/~smaloy/MicrobialGenetics/topics/in-vitro-genetics/PCR.html. "Polymerase Chain Reaction (PCR)" Biology Animation Library :: DNA Learning Center." Dnalc.Org, https://www.dnalc.org/resources/animations/pcr.html. "Polymerase Chain Reaction (PCR)." Khan Academy, 2018, https://www.khanacademy.org/science/biology/biotech-dna-technology/dna-sequencing-pcr-electrophoresis/a/polymerase-chain-reaction-pcr. "Polymerase Chain Reaction (PCR)." Biology-Pages.Info, 2014, http://www.biology-pages.info/P/PCR.html. "Openstax CNX." Cnx.Org, 2016, https://cnx.org/contents/[email protected]:exg4e4AU@7/Biotechnology. "Polymerase Chain Reaction (PCR)." Ncbi.Nlm.Nih.Gov, 2017, https://www.ncbi.nlm.nih.gov/probe/docs/techpcr/. "Polymerase Chain Reaction / PCR | Learn Science At Scitable." Nature.Com, https://www.nature.com/scitable/definition/polymerase-chain-reaction-pcr-110. "The Role Of Taq Polymerase In PCR." Sciencing.Com, 2017, https://sciencing.com/role-taq-polymerase-pcr-7298417.html. Phillips, Theresa. "Visualizing DNA." The Balance, 2018, https://www.thebalance.com/visualizing-dna-375499. "Electrophoresis." Bioinformatics.Nl, http://www.bioinformatics.nl/molbi/SimpleCloningLab/electrophoresis.htm.
"Thermus Aquaticus." En.Wikipedia.Org, https://en.wikipedia.org/wiki/Thermus_aquaticus#/media/File:Thermus_aquaticus.JPG.
0 Comments
Your comment will be posted after it is approved.
Leave a Reply. |
AuthorHello, I am Elaine. A junior at Holland Hall School. I am passionate about learning genomics and genome editing. In this blog page, you will see lab reports, personal opinion pieces, and some essays on biology! Archives
April 2018
Categories
|























 RSS Feed
RSS Feed
